# Load required packages
library(tidyverse)
library(broom)
library(rstatix)Friedman Test Analysis
Friedman Test Analysis
Friedman’s test is a non-parametric statistical test used to detect differences in treatments across multiple test attempts. It is often used when the assumptions of ANOVA are not met, particularly the assumption of normality. The test is applicable for repeated measures, or matched groups, making it useful for situations where the same subjects are subjected to different treatments.
Friedman’s test ranks the data points within each block (or subject) separately, and then analyzes these ranks to see if the mean ranks differ between the groups and conditions. If the test shows significant differences, this suggests that at least one of the treatments differs from the others. Because it is non-parametric, it does not assume the normal distribution of data, which makes it robust for skewed or ordinal data.
Hypotheses
H₀ (Null Hypothesis): There are no significant differences in weight outcomes between the three diets
H₁ (Alternative Hypothesis): There are significant differences in weight outcomes between at least two diets
# Create the dataset
Diet_A = c(75, 68, 80, 72, 85, 70, 82, 78, 75, 83)
Diet_B = c(82, 70, 85, 78, 88, 75, 85, 80, 79, 87)
Diet_C = c(78, 65, 82, 75, 84, 72, 80, 76, 77, 84)
data <- tibble(
subjid = rep(1:10, 3),
diet = rep(c("A", "B", "C"), each = 10),
weight = c(Diet_A, Diet_B, Diet_C)
)Base R {stats}
To run a Friedman’s test in R you can use the {stats} package. This will return the chi-squared test statistic and p-value.
# Perform Friedman test
friedman_test <- stats::friedman.test(weight ~ diet | subjid, data = data)
friedman_test |>
broom::tidy()# A tibble: 1 × 4
statistic p.value parameter method
<dbl> <dbl> <dbl> <chr>
1 15.2 0.000500 2 Friedman rank sum test{rstatix}
Alternatively, you can use the {rstatix} package. While these packages give the same results, the {rstatix} results come as a tibble we can easily use.
test <- data |>
rstatix::friedman_test(weight ~ diet | subjid)
test# A tibble: 1 × 6
.y. n statistic df p method
* <chr> <int> <dbl> <dbl> <dbl> <chr>
1 weight 10 15.2 2 0.000500 Friedman test# Create boxplot
ggplot(data, aes(x = diet, y = weight, fill = diet)) +
geom_boxplot() +
theme_minimal() +
labs(
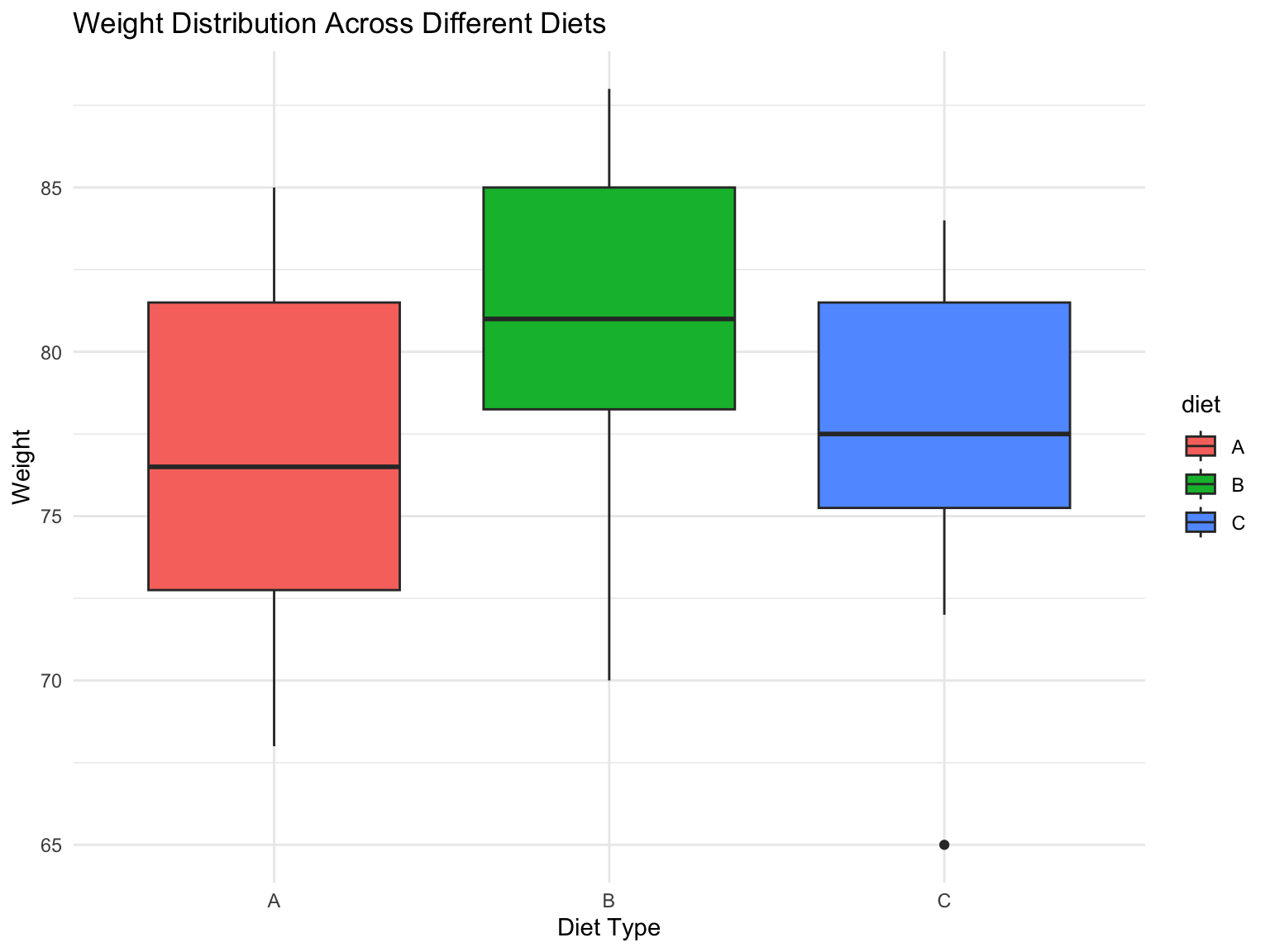
title = "Weight Distribution Across Different Diets",
x = "Diet Type",
y = "Weight"
)
Conclusion
Based on the analysis:
- Statistical Test Results:
- The Friedman test yielded a p-value of 5.0045143^{-4}
[If p < 0.05, we reject the null hypothesis
If p > 0.05, we fail to reject the null hypothesis]
- Visual Analysis:
From the boxplot, Diet B shows the highest median weight
Diet B also appears to have the highest overall weight distribution
Diet A and Diet C show similar distributions but lower than Diet B
- Interpretation:
If the goal is weight gain: Diet B appears most effective
If the goal is weight maintenance: Diet A or C might be more suitable
However, individual responses vary, as shown by the overlapping distributions
Reference
Cite all sources and references used in the analysis.
─ Session info ───────────────────────────────────────────────────────────────
setting value
version R version 4.5.2 (2025-10-31)
os macOS Tahoe 26.3
system aarch64, darwin20
ui X11
language (EN)
collate en_US.UTF-8
ctype en_US.UTF-8
tz Europe/London
date 2026-02-23
pandoc 3.6.3 @ /Applications/Positron.app/Contents/Resources/app/quarto/bin/tools/aarch64/ (via rmarkdown)
quarto 1.8.27 @ /Applications/Positron.app/Contents/Resources/app/quarto/bin/quarto
─ Packages ───────────────────────────────────────────────────────────────────
! package * version date (UTC) lib source
P abind 1.4-8 2024-09-12 [?] RSPM
askpass 1.2.1 2024-10-04 [1] RSPM
P backports 1.5.0 2024-05-23 [?] RSPM
base64enc 0.1-6 2026-02-02 [1] RSPM
bit 4.6.0 2025-03-06 [1] RSPM
bit64 4.6.0-1 2025-01-16 [1] RSPM
blob 1.3.0 2026-01-14 [1] RSPM
boot 1.3-32 2025-08-29 [2] CRAN (R 4.5.2)
P broom * 1.0.12 2026-01-27 [?] RSPM
bslib 0.10.0 2026-01-26 [1] RSPM
cachem 1.1.0 2024-05-16 [1] RSPM
callr 3.7.6 2024-03-25 [1] RSPM
P car 3.1-5 2026-02-03 [?] RSPM
P carData 3.0-6 2026-01-30 [?] RSPM
cellranger 1.1.0 2016-07-27 [1] RSPM
P cli 3.6.5 2025-04-23 [?] RSPM
clipr 0.8.0 2022-02-22 [1] RSPM
colorspace 2.1-2 2025-09-22 [1] RSPM
conflicted 1.2.0 2023-02-01 [1] RSPM
corrplot 0.95 2024-10-14 [1] RSPM
cowplot 1.2.0 2025-07-07 [1] RSPM
cpp11 0.5.3 2026-01-20 [1] RSPM
crayon 1.5.3 2024-06-20 [1] RSPM
curl 7.0.0 2025-08-19 [1] RSPM
data.table 1.18.2.1 2026-01-27 [1] RSPM
DBI 1.2.3 2024-06-02 [1] RSPM
dbplyr 2.5.2 2026-02-13 [1] RSPM
Deriv 4.2.0 2025-06-20 [1] RSPM
P digest 0.6.39 2025-11-19 [?] RSPM
doBy 4.7.1 2025-12-02 [1] RSPM
P dplyr * 1.2.0 2026-02-03 [?] RSPM
dtplyr 1.3.3 2026-02-11 [1] RSPM
P evaluate 1.0.5 2025-08-27 [?] RSPM
P farver 2.1.2 2024-05-13 [?] RSPM
P fastmap 1.2.0 2024-05-15 [?] RSPM
fontawesome 0.5.3 2024-11-16 [1] RSPM
P forcats * 1.0.1 2025-09-25 [?] RSPM
forecast 9.0.1 2026-02-14 [1] RSPM
P Formula 1.2-5 2023-02-24 [?] RSPM
fracdiff 1.5-3 2024-02-01 [1] RSPM
fs 1.6.6 2025-04-12 [1] RSPM
gargle 1.6.1 2026-01-29 [1] RSPM
P generics 0.1.4 2025-05-09 [?] RSPM
P ggplot2 * 4.0.2 2026-02-03 [?] RSPM
P glue 1.8.0 2024-09-30 [?] RSPM
googledrive 2.1.2 2025-09-10 [1] RSPM
googlesheets4 1.1.2 2025-09-03 [1] RSPM
P gtable 0.3.6 2024-10-25 [?] RSPM
haven 2.5.5 2025-05-30 [1] RSPM
highr 0.11 2024-05-26 [1] RSPM
P hms 1.1.4 2025-10-17 [?] RSPM
P htmltools 0.5.9 2025-12-04 [?] RSPM
httr 1.4.8 2026-02-13 [1] RSPM
ids 1.0.1 2017-05-31 [1] RSPM
isoband 0.3.0 2025-12-07 [1] RSPM
jquerylib 0.1.4 2021-04-26 [1] RSPM
P jsonlite 2.0.0 2025-03-27 [?] RSPM
P knitr 1.51 2025-12-20 [?] RSPM
P labeling 0.4.3 2023-08-29 [?] RSPM
lattice 0.22-7 2025-04-02 [2] CRAN (R 4.5.2)
P lifecycle 1.0.5 2026-01-08 [?] RSPM
lme4 1.1-38 2025-12-02 [1] RSPM
lmtest 0.9-40 2022-03-21 [1] RSPM
P lubridate * 1.9.5 2026-02-04 [?] RSPM
P magrittr 2.0.4 2025-09-12 [?] RSPM
MASS 7.3-65 2025-02-28 [2] CRAN (R 4.5.2)
Matrix 1.7-4 2025-08-28 [2] CRAN (R 4.5.2)
MatrixModels 0.5-4 2025-03-26 [1] RSPM
memoise 2.0.1 2021-11-26 [1] RSPM
mgcv 1.9-3 2025-04-04 [2] CRAN (R 4.5.2)
microbenchmark 1.5.0 2024-09-04 [1] RSPM
mime 0.13 2025-03-17 [1] RSPM
minqa 1.2.8 2024-08-17 [1] RSPM
modelr 0.1.11 2023-03-22 [1] RSPM
nlme 3.1-168 2025-03-31 [2] CRAN (R 4.5.2)
nloptr 2.2.1 2025-03-17 [1] RSPM
nnet 7.3-20 2025-01-01 [2] CRAN (R 4.5.2)
numDeriv 2016.8-1.1 2019-06-06 [1] RSPM
openssl 2.3.4 2025-09-30 [1] RSPM
pbkrtest 0.5.5 2025-07-18 [1] RSPM
P pillar 1.11.1 2025-09-17 [?] RSPM
P pkgconfig 2.0.3 2019-09-22 [?] RSPM
prettyunits 1.2.0 2023-09-24 [1] RSPM
processx 3.8.6 2025-02-21 [1] RSPM
progress 1.2.3 2023-12-06 [1] RSPM
ps 1.9.1 2025-04-12 [1] RSPM
P purrr * 1.2.1 2026-01-09 [?] RSPM
quantreg 6.1 2025-03-10 [1] RSPM
P R6 2.6.1 2025-02-15 [?] RSPM
ragg 1.5.0 2025-09-02 [1] RSPM
rappdirs 0.3.4 2026-01-17 [1] RSPM
rbibutils 2.4.1 2026-01-21 [1] RSPM
P RColorBrewer 1.1-3 2022-04-03 [?] RSPM
Rcpp 1.1.1 2026-01-10 [1] RSPM
RcppArmadillo 15.2.3-1 2025-12-17 [1] RSPM
RcppEigen 0.3.4.0.2 2024-08-24 [1] RSPM
Rdpack 2.6.6 2026-02-08 [1] RSPM
P readr * 2.1.6 2025-11-14 [?] RSPM
readxl 1.4.5 2025-03-07 [1] RSPM
reformulas 0.4.4 2026-02-02 [1] RSPM
rematch 2.0.0 2023-08-30 [1] RSPM
rematch2 2.1.2 2020-05-01 [1] RSPM
reprex 2.1.1 2024-07-06 [1] RSPM
P rlang 1.1.7 2026-01-09 [?] RSPM
P rmarkdown 2.30 2025-09-28 [?] RSPM
P rstatix * 0.7.3 2025-10-18 [?] RSPM
rstudioapi 0.18.0 2026-01-16 [1] RSPM
rvest 1.0.5 2025-08-29 [1] RSPM
P S7 0.2.1 2025-11-14 [?] RSPM
sass 0.4.10 2025-04-11 [1] RSPM
P scales 1.4.0 2025-04-24 [?] RSPM
selectr 0.5-1 2025-12-17 [1] RSPM
SparseM 1.84-2 2024-07-17 [1] RSPM
P stringi 1.8.7 2025-03-27 [?] RSPM
P stringr * 1.6.0 2025-11-04 [?] RSPM
survival 3.8-3 2024-12-17 [2] CRAN (R 4.5.2)
sys 3.4.3 2024-10-04 [1] RSPM
systemfonts 1.3.1 2025-10-01 [1] RSPM
textshaping 1.0.4 2025-10-10 [1] RSPM
P tibble * 3.3.1 2026-01-11 [?] RSPM
P tidyr * 1.3.2 2025-12-19 [?] RSPM
P tidyselect 1.2.1 2024-03-11 [?] RSPM
P tidyverse * 2.0.0 2023-02-22 [?] RSPM
P timechange 0.4.0 2026-01-29 [?] RSPM
timeDate 4052.112 2026-01-28 [1] RSPM
tinytex 0.58 2025-11-19 [1] RSPM
P tzdb 0.5.0 2025-03-15 [?] RSPM
urca 1.3-4 2024-05-27 [1] RSPM
P utf8 1.2.6 2025-06-08 [?] RSPM
uuid 1.2-2 2026-01-23 [1] RSPM
P vctrs 0.7.1 2026-01-23 [?] RSPM
viridisLite 0.4.3 2026-02-04 [1] RSPM
vroom 1.7.0 2026-01-27 [1] RSPM
P withr 3.0.2 2024-10-28 [?] RSPM
P xfun 0.56 2026-01-18 [?] RSPM
xml2 1.5.2 2026-01-17 [1] RSPM
P yaml 2.3.12 2025-12-10 [?] RSPM
zoo 1.8-15 2025-12-15 [1] RSPM
[1] /Users/christinafillmore/Documents/GitHub/CAMIS/renv/library/macos/R-4.5/aarch64-apple-darwin20
[2] /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/library
* ── Packages attached to the search path.
P ── Loaded and on-disk path mismatch.
──────────────────────────────────────────────────────────────────────────────